Oil and natural gas transportation through pipelines—both onshore and offshore—is considered a safer alternative on a ton-mile basis to hauling fuel by tanker trucks, freight trains, or marine carriers [1]. Even so, the transportation of liquid or gas-phase hydrocarbons over long distances can lead to accidental releases via pipeline ruptures or leaks. The number and severity of incidents related to leaking pipelines, and the spills associated with them, have increased substantially over the last three decades (see Figure 1) [2].
Although safer than the alternatives for fuel transportation, pipeline networks require thorough maintenance and upkeep to remain operable. Pipeline ruptures can result in significant economic, environmental, and energy-security consequences. The energy system of the United States is one of 16 infrastructure sectors identified by the Department of Homeland Security (DHS) as “critical” to the nation’s economic security, public health, safety, and defense [3,4]. The department’s sector-specific plan for critical energy infrastructure notes that pipelines face two major risks: operational hazards caused by blowouts, spills, and personal injury; and interruptions in energy supplies due to damaged or inoperable hydrocarbon pipeline infrastructure [4].
Technologies that enable operators to detect small leaks before a rupture or major release incident occurs can significantly reduce these risks. The early detection of small leaks allows remedial action to ensure continued pipeline integrity [5]. The need for a new class of hydrocarbon sensors is especially pressing, because sensor types currently in use for onshore pipelines in the United States often fail to detect releases. Only 22 percent of the releases that occurred between 2010 and 2016 were detected by advanced pipeline sensor systems [6].
Current State of the Art
Well-known methods for leak detection in onshore pipelines include acoustic monitoring, optical sensor gas sampling, soil monitoring, flow monitoring, magnetic flux leakage, and dynamic model-based methods [9]. “Pig” (or pipeline-inspection-gadget) technology is used to detect stress corrosion cracking, uniform or general corrosion, and pitting corrosion—including gouges, dents, anomalous weld seams, longitudinal cracks, longitudinal grooves, and other faults in the pipeline wall [10]. In-pipe robots used for early detection include: pig type, wheel type, caterpillar type, walking type, inchworm type, screw type, and wall-press. Another monitoring technique involves the use of unmanned aerial vehicles (or autonomous underwater vehicle) for pipeline and flowline surveillance [10]. The major drawbacks of the inspection robots presently used in the oil and gas industry center on insufficiently high levels of desired autonomy, robustness, and dependability.
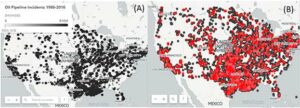
Figure 1. (A) Plot of economic damages caused by significant oil and natural gas pipeline-associated incidents and (B) number of resulting fatalities, 1986–2016. These incidents have resulted in 548 deaths, 2,576 injuries, and more than $8.5 billion in financial damages over
this time period [2].
A variety of sensor technologies are capable of remotely detecting hydrocarbon releases from offshore pipelines using techniques that rely on the relative adsorption of visible, infrared, and ultraviolet irradiation by an oil slick [7,8]. Adsorption intensity is related to the thickness of the oil layer on the water surface. Radar technologies, as well as microwave radiometers, laser fluoro-sensors, and laser-acoustic approaches, have also been used as oil thickness sensors. These sensors can provide information on the location and spread of an oil spill; its thickness distribution,
which allows for a volumetric estimation; and the type and quality of the oil (i.e., its API gravity and H2S content), which aids in assessing environmental damage and take appropriate actions for clean-up [7, 8].
When employing these techniques, one important consideration is the cost of data acquisition and analysis, which includes measurement equipment, operational and personnel costs, and data collection. Remote detection sensors are typically deployed in air-borne mode, adding significant cost to operations. Moreover, with regard to offshore pipeline rupture sensing, their data output is influenced by a plethora of conditions related to water flow, water quality, metocean and weather conditions. While the deployment of these technologies is significant, more robust and reliable sensor technologies with improved feasibility and reliability are desired for future applications [10].
Additionally, the sensor technologies outlined above generally do not provide information to implement a proactive strategy for the prevention of hydrocarbon releases. Most provide post-incident detection and notification, and those capable of early detection of deficiencies in pipeline integrity (e.g., pigs), do not deliver continuous monitoring. In-situ microbial sensors may have a unique ability to achieve this important outcome.
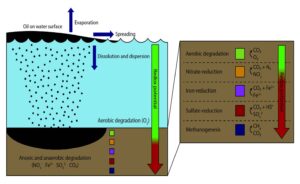
Figure 2. Fate and transport of oil leakages or spills in the marine environment and biodegradation (redox) potentials [13].
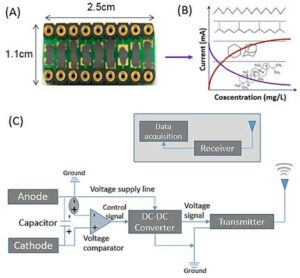
Figure 3. (A) a centimeter-scale microbial electrochemical sensor (microbial–microalgae); (B) evaluation of sensor response to hydrocarbon and petroleum compounds; and (C) a sensor platform for signal transmission, data acquisition, and signal processing.
A Novel Microbial Sensor
Early detection of small liquid or gas hydrocarbon leaks or spills could lessen the likelihood of a major release incident. However, organic contaminant sensors are difficult to employ under certain settings due to material- or function-related constraints usually associated with background concentrations, mass-transfer, and detection limitations. In addition, environmental variations in temperature, barometric pressure, and relative humidity play an important role in subsurface environments [12]. Our research focuses on developing a self-sustained microbial (living-cell) sensor platform suitable for harsh or remote environments, such as those found at deep-water subsea infrastructure, onshore surface and subsurface pipelines, and hydrocarbon processing plants.
A microbial electrochemical cell (biosensor) was developed in order to harness the benefits of biodiversity in microbial populations and in their metabolic functions, thus creating a variety of electrochemical cells suitable for applications in varying environmental settings. The biosensor was created using exoelectrogenic bacteria that survives on organic matter in an anode and a phototrophic, autotrophic, or sulfur-reducing bacterial consortium in the cathode—enabling continuous transfer of electrons from one compartment (anode) to the other (cathode) [14].
This arrangement generates a voltage across a resistor which can be set or tuned to identify possible contamination by hydrocarbon compounds. The increased availability of organic compounds within the microbial biosensor cell will generate high voltage spikes. A variety of hydrocarbons and organic compounds (e.g., crude oil, natural gas, refined gasoline) can be used as substrates in laboratory studies to understand voltage generation potential and to optimize and stabilize sensor operations for different target petroleum products [15].
As shown in Figure 2, the oxidation-reduction potentials present in the marine, sediment, and above-ground onshore environments provide numerous opportunities for utilizing natural processes to simultaneously detect and communicate hydrocarbon releases. Microbial fuel cells (MFCs) have been used to run and power remotely-operated biological oxygen demand sensors [17]. Sediment-based MFCs have also been used for powering a meteorological buoy, wireless temperature sensors [18], and other environmental sensors [19]. MFCs have also functioned as power sources for remote sensors, digital wrist watches [20], light-emitting diodes for internal lighting [21], and basic phone/smart phone charging [22].
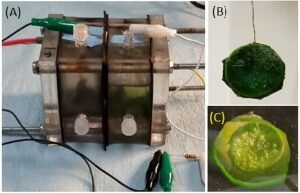

Figure 4. (A) A microbial biosensor with mixed bacterial consortium in the anode chamber and a microalgae biocathode in the cathode chamber; (B) a “mutag BioChip 25™” with a carbon felt electrode used to concentrate microalgae biofilm; and (C) in-situ oxygen generation
by microalgae cells residing on the electrode.
To overcome the restricted lifetime of batteries used in traditional, non-biological sensors, microbial sensor technology can be used as a telemetry system that transmits signals to remote receivers. To develop an effective and reliable biological sensor based on this technology, appropriate anodic and cathodic reactions should be combined. This enables the technology to be used for detection of organic pollutants, such as oil spills, and the communication of such. Several years of operational experience have been reported for other applications, including in biochemical oxygen demand sensors [17]. Because microbial systems have already proved useful for remediating toxic substances—such as phenols and petroleum compounds [23]—they should be explored for novel applications in extreme settings.
Our research approach is depicted in Figure 3. Three different types of immobilized microbial sensors will be developed to adapt to deep marine saline and sediment environments as well as surface and subsurface environments. These microbial sensors use exoelectrogenic bacteria (anode) as well as anammox and microalgae (namely, Chlorella Vulgaris sp. [cathode]) as anode and cathode consortia [24-26]. The specific objectives are:
- to develop centimeter-scale microbial electrochemical cells (see Figure 3a) suitable for marine/soil/surface environmental settings using various combinations of exoelectrogenic, autotrophic,and phototrophic microbial consortia;
- to evaluate the performance of the biosensors in terms of voltage generation potentials through the use of hydrocarbon- and petroleum-based compounds (see Figure 3b);
- to optimize biosensor performance by varying environmental settings, including pH, water depth, temperature, and substrate/nutrient concentrations; and
- to develop a sensor platform using voltage- to-acoustic signal transfer, data acquisition, and signal processing (see Figure 3c).
Our microbial biosensor prototype, shown in Figure 4, was used to carry out a series of experiments that build upon the findings of our previous research on biofilm development and experimental procedures [27, 28]. After a stable biofilm had been produced in the biosensor and a stable voltage achieved (under the reference physiological settings), the unit was subjected to various concentrations of gasoline (by volume/volume) in both anode and cathode chambers in order to mimic real-world conditions of a hydrocarbon release into the environment.
The microbial sensor’s response to various gasoline concentrations during more than 1,000 hours of operation can be seen in Figure 5. The first six experiments (represented by the blue columns in Figure 5) were performed to allow for the start-up and establishment of a stable biofilm in both anode and cathode chambers; achievement of this is confirmed by the voltage generated by the cell. The maximum voltage (peak) produced by the cell was 0.1201 V at a chemical oxygen demand (COD) concentration of 500 mg/L. Three repetitive cycles were conducted which yielded an average response value of 0.1151 V. The microbial sensor was then subjected to a very low concentration of gasoline with 10 percent ethanol (0.04% v/v, shown in purple in Figure 5). The response of the microbial sensor remained the same, demonstrating that it was not sensitive to a very low level of gasoline concentration.
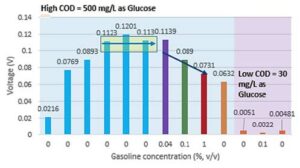
Figure 5. Response of a microbial-microalgae sensor with respect to the varying COD (30 mg/L and 500 mg/L) and Gasoline concentrations (0-1% v/v).
In the next experiment, the gasoline concentration was increased by 2.5 times (i.e., 0.1% v/v) from the previous experiment. The response of the sensor was significantly affected by the gasoline, resulting in a 23 percent reduction in the response (shown in green in Figure 5). When the concentration was increased by 10 times (i.e., from 0.1% to 1% v/v ratio), the response was decreased by 37% from its original gasoline test level (shown in red in Figure 5). Following these experiments, the microbial sensor was returned to its original condition (without the addition of gasoline). Overall, a linear relationship was observed between the voltage response and the gasoline concentrations of 0% and 0.04%. An exponential relationship was observed for the voltage response
between 0.04% and 1.0% v/v.
Another set of experiments was conducted at low COD concentrations to understand the response of the microbial sensor under low substrate conditions (shown in purple in Figure 5). There was a 57 percent reduction in voltage at 0.1% v/v gasoline addition at a COD concentration of 30 mg/L. The sensor was able to recover its voltage response when the media was replaced with a low COD wastewater without a gasoline addition. A matrix of experiments is required to further develop response relationships to variations of hydrocarbon compounds. For example, gasoline with 10 percent ethanol, which contains a mixture of long chain hydrocarbons, was used as a model compound in this research.
Diesel has a different composition, however, and a microbial sensor will exhibit a different response when exposed to it. Therefore, a series of experiments and optimization studies are required to develop a robust microbial sensor that would produce stable and reliable responses when exposed to hydrocarbon releases from a wide array of fuel types.
Future Perspective
The proposed microbial electrochemical cell (biosensor) platform produces a voltage spike or sink in response to a certain environment. Moreover, sensors using different types of microbes can be used to detect an array of target molecules. Researchers have identified several microorganism groups that readily degrade hydrocarbons, especially in aquatic and marine environments [29, 30].
Recent research suggests that Alcanivorax borkumensis marine bacteria in particular are specialized in utilizing alkanes for growth. Several other microorganisms belonging to bacteria, yeast, and fungi groups have also been identified for their potential to degrade crude oils and hydrocarbons [29]. Several members of these communities could be employed in the microbial sensor platform to develop a substantial and traceable response to the availability of hydrocarbons.
While this is an attractive approach, little knowledge exists on this microbial community structure and their function in environments exposed to oil, gasoline, or natural gas spills and leaks [31]. Future research efforts should focus on developing a deeper understanding of the functional genes responsible for oil degradation in marine and soil environments. If this knowledge is developed comprehensively, midstream petroleum companies, pipeline network operators, and government and military leaders in the homeland defense community may be able to quickly, accurately, and precisely respond to small hydrocarbon releases before a rupture threatens economic, environmental, and energy security.
Acknowledgement
The authors would like to thank Multi Umwelttechnologie AG of Germany for providing the Mutag BioChip25™ samples used in this study.
References
1. Furchtgott-Roth, D. (2013, June 6). Pipelines are safest for transportation of oil and gas. Manhattan Institute for Policy Research Issue Brief, 23, 1–10. Retrieved from https:// www.manhattan-institute.org/pdf/ib_23.pdf
2. Joseph, G. (2016, November 30). 30 years of oil and gas pipeline accidents, mapped. CityLab. Retrieved from https://www.citylab. com/environment/2016/11/30-years-of-pipeline-accidents-mapped/509066/
3. U.S. Department of Homeland Security. (2013, December). National Infrastructure Protection Plan (NIPP) 2013: Partnering for critical infrastructure security and resilience. Retrieved from https://www.dhs.gov/sites/default/files/publications/national-infrastructure-protection-plan-2013-508.pdf
4. U.S. Department of Homeland Security. (2015). Energy Sector-Specific Plan. Retrieved from https://www.dhs.gov/sites/default/files/publications/nipp-ssp-energy-2015-508.pdf
5. Canada’s Oil and Natural Gas Producers. (2018, May 26). Best management practice: Pipeline leak detection programs. Retrieved from https://www.capp.ca/~/media/capp/customer-portal/publications/310502.pdf?-modified=20180526153412
6. Renshaw, J., & Kumar, D. K. (2016, September 30). Technology designed to detect U.S. energy pipeline leaks often fails. Reuters. Retrieved from https://www.reuters.com/article/us-usa-pipelines-colonial-analysis/technology-designed-to-detect-u-s-energypipeline-leaks-often-fails-idUSKCN1200FQ
7. Grüner, K., Reuter, R., & Smid, H. (1991). A new sensor system for airborne measurements of maritime pollution and of hydrographic parameters. GeoJournal, 24(1),103–117. doi:10.1007/bf00213062
8. Jha, M., Levy, J., & Gao, Y. (2008). Advances in remote sensing for oil spill disaster management: State-of-the-art sensors technology for oil spill surveillance. Sensors, 8(1), 236–255. doi:10.3390/s8010236
9. Liu, M., Zang, S., & Zhou, D. (2005). Fast leak detection and location of gas pipelines based on an adaptive particle filter. International Journal of Applied Mathematics and Computer Science, 15(4), 541–550. Retrieved from http://matwbn.icm.edu.pl/ksiazki/amc/amc15/amc15411.pdf
10. Shukla, A., & Karki, H. (2016). Application of robotics in onshore oil and gas industry—A review Part I. Robotics and Autonomous Systems, 75, 490–507. doi:10.1016/j.robot.
2015.09.012
11. Albiez, J., Hildebrand, M., Vogele, T., Joyeux, S., & Kirchner, F. (2011, January 1). Robust Robots for Arctic Exploration. Offshore Technology Conference. doi:10.4043/22120-MS
12. Ho, C. K., & Hughes, R. C. (2002). In-situ chemiresistor sensor package for real-time detection of volatile organic compounds in soil and groundwater. Sensors, 2(1), 23–34.
doi:10.3390/s20100023
13. Daghio, M., Aulenta, F., Vaiopoulou, E., Franzetti, A., Arends, J. B., Sherry, A., … & Rabaey, K. (2017). Electrobioremediation of oil spills. Water Research, 114, 351–370.
doi:10.1016/j.watres.2017.02.030
14. Gude, V. G. (2016). Wastewater treatment in microbial fuel cells–an overview. Journal of Cleaner Production, 122, 287–307. doi:10.1016/j.jclepro.2016.07.048
15. Tecon, R., der Meer, V., & Roelof, J. (2008).Bacterial biosensors for measuring availability of environmental pollutants. Sensors, 8(7), 4062–4080. doi:10.3390/s8074062
16. Du, Z., Li, H., & Gu, T. (2007). A state of the art review on microbial fuel cells: a promising technology for wastewater treatment and bioenergy. Biotechnology Advances, 25(5), 464–482. doi:10.1016/j.biotechadv.2007.05.004
17. Chang, I. S., Moon, H., Jang, J. K., & Kim, B. H. (2005). Improvement of a microbial fuel cell performance as a BOD sensor using respiratory inhibitors. Biosensors and Bioelectronics,
20(9), 1856–1859. doi:10.1016/j. bios.2004.06.003
18. Tender, L. M., Gray, S. A., Groveman, E., Lowy, D. A., Kauffman, P., Melhado, J., … & Dobarro, J. (2008). The first demonstration of a microbial fuel cell as a viable power
supply: powering a meteorological buoy. Journal of Power Sources, 179(2), 571–575. doi:10.1016/j.jpowsour.2007.12.123
19. Donovan, C., Dewan, A., Peng, H., Heo, D., & Beyenal, H. (2011). Power management system for a 2.5 W remote sensor powered by a sediment microbial fuel cell. Journal of Power Sources, 196(3), 1171–1177. doi:10.1016/j.jpowsour.2010.08.099
20. Papaharalabos, G., Greenman, J., Melhuish, C., Santoro, C., Cristiani, P., Li, B., & Ieropoulos, I. (2013). Increased power output from micro porous layer (MPL) cathode microbial fuel cells (MFC). International Journal of Hydrogen Energy, 38(26), 11552–11558. doi:10.1016/j.ijhydene.2013.05.138
21. Ieropoulos, I. A., Ledezma, P., Stinchcombe, A., Papaharalabos, G., Melhuish, C., & Greenman, J. (2013). Waste to real energy: the first MFC powered mobile phone. Physical Chemistry Chemical Physics, 15(37), 15312–15316. doi:10.1039/C3CP52889H
22. Walter, X. A., Stinchcombe, A., Greenman, J., & Ieropoulos, I. (2017). Urine transduction to usable energy: a modular MFC approach for smartphone and remote system charging. Applied Energy, 192, 575–581. doi:10.1016/j.apenergy.2016.06.006
23. Morris, J. M., & Jin, S. (2007). Feasibility of using microbial fuel cell technology for bioremediation of hydrocarbons in groundwater. Journal of Environmental Science and Health, Part A, 43(1), 18–23. doi:10.1080/10934520701750389
24. Kokabian, B., Smith, R., Brooks, J. P., & Gude, V. G. (2018). Bioelectricity production in photosynthetic microbial desalination cells under different flow configurations. Journal of Industrial and Engineering Chemistry, 58, 131–139. doi:10.1016/j.jiec.2017.09.017
25. Kokabian, B., Ghimire, U., & Gude, V. G. (2018). Water deionization with renewable energy production in microalgae- microbial desalination process. Renewable Energy, 122, 354–361. doi:10.1016/j.renene.2018.01.061
26. Kokabian, B., Gude, V. G., Smith, R., & Brooks, J. P. (2018). Evaluation of anammox biocathode in microbial desalination and wastewater treatment. Chemical Engineering Journal, 342, 410–419. doi:10.1016/j.cej.2018.02.088
27. Kokabian, B., & Gude, V. G. (2013). Photosynthetic microbial desalination cells (PMDCs) for clean energy, water and biomass production. Environmental Science: Processes & Impacts, 15(12), 2178–2185. doi:10.1039/c3em00415e
28. Kokabian, B., & Gude, V. G. (2015). Sustainable photosynthetic biocathode in microbial desalination cells. Chemical Engineering Journal, 262, 958–965. doi:10.1016/j.cej.2014.10.048
29. Hazen, T. C., Prince, R. C., & Mahmoudi, N. (2016). Marine oil biodegradation. Environmental Science & Technology, 50(5), 2121–2129. doi:10.1021/acs.est.5b03333
30. Hazen, T. C., Dubinsky, E. A., DeSantis, T. Z., Andersen, G. L., Piceno, Y. M., Singh, N., … & Stringfellow, W. T. (2010). Deep-sea oil plume enriches indigenous oil-degrading bacteria. Science, 330(6001), 204–208. doi:10.1126/science.1195979
31. Mason, O. U., Hazen, T. C., Borglin, S., Chain, P. S., Dubinsky, E. A., Fortney, J. L.,… & Mackelprang, R. (2012). Metagenome, metatranscriptome and single-cell sequencing reveal microbial response to Deepwater Horizon oil spill. The ISME Journal, 6(9), 1715. doi:10.1038/ismej.2012.59


